For practicing bioinformatics techniques in class, you can use any data set that’s not beyond the capacity of your computer. In practice that means that you probably want to use a bacterial NGS data set.
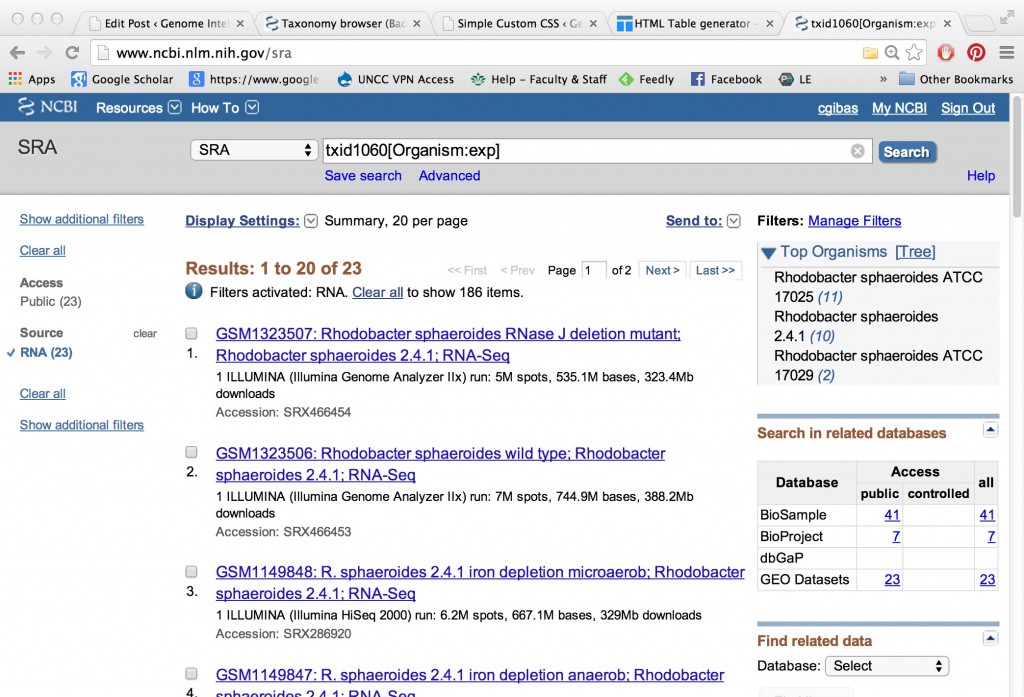
It’s possible to search all of NCBI to find which genomes have datasets available in the short read archive. (It’s not quite as possible to easily catalog them for a meta-study or download them en masse.) Try going in through the NCBI Taxonomy Browser and clicking the “SRA Experiments” checkbox to get a taxonomy view labeled with links to experiments in the SRA.
You can narrow things down by using a species or genus name that you are familiar with, for instance here I started at “Rhodobacter” because I know there is a whole bunch of published Illumina and Pac Bio data from the Broad Institute from their comparative assembly studies for development of the ALLPATHS assembler.
Genomes to assemble
A genome with paired end Illumina data only (V. vulnificus JY1305, which we published here).
The ION Torrent single-end chloroplast genome sequencing data generated in BINF 6350 can be assembled using some assemblers that are optimized for mid-length reads (i.e. not Velvet; something like MIRA or Newbler — or CLC, of course — is appropriate).
A Staph aureus MW2 genome with multiple insert lengths (paired end and mate pair).
Multi-strain comparision/variant detection
The chloroplast genome sequencing data generated in your BINF 6350 class can be used to practice read mapping and variant detection analysis.
RNA-Seq
We’re about to deposit a Vibrio vulnificus RNA-Seq data set (Illumina) that will be excellent for use in class, though it’s still unavoidably somewhat big. Replicate data for four Vibrio vulnificus strains in human serum vs. artificial seawater are available. For 2014’s class you still need to check this data out from me on thumb drives.
Here’s a small comparative transcriptomics data set in Rhodobacter sphaeroides, wild type and with RNAseJ knocked out.
Here is a Synechocystis transcriptome data set under 10 different conditions (unreplicated).
ChIP-Seq
Small ChIP-Seq data set for sigma70 in WT and HNS mutant E. coli K12.
Was this helpful?
0 /
0